Abstract
Key message
The study on the GmDWF1-deficient mutant dwf1 showed that GmDWF1 plays a crucial role in determining soybean plant height and yield by influencing the biosynthesis of brassinosteroids.
Abstract
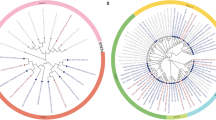
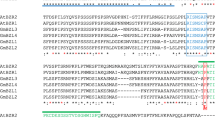
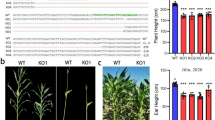
Soybean has not adopted the Green Revolution, such as reduced height for increased planting density, which have proven beneficial for cereal crops. Our research identified the soybean genes GmDWF1a and GmDWF1b, homologous to Arabidopsis AtDWF1, and found that they are widely expressed, especially in leaves, and linked to the cellular transport system, predominantly within the endoplasmic reticulum and intracellular vesicles. These genes are essential for the synthesis of brassinosteroids (BR). Single mutants of GmDWF1a and GmDWF1b, as well as double mutants of both genes generated through CRISPR/Cas9 genome editing, exhibit a dwarf phenotype. The single-gene mutant exhibits moderate dwarfism, while the double mutant shows more pronounced dwarfism. Despite the reduced stature, all types of mutants preserve their node count. Notably, field tests have shown that the single GmDWF1a mutant produced significantly more pods than wild-type plants. Spraying exogenous brassinolide (BL) can compensate for the loss in plant height induced by the decrease in endogenous BRs. Comparing transcriptome analyses of the GmDWF1a mutant and wild-type plants revealed a significant impact on the expression of many genes that influence soybean growth. Identifying the GmDWF1a and GmDWF1b genes could aid in the development of compact, densely planted soybean varieties, potentially boosting productivity.






Similar content being viewed by others
Data availability
The data sets generated during the current study are available from the corresponding author on reasonable request. RNA-Seq’s raw data has been published at WEBLINK https://www.ncbi.nlm.nih.gov/sra/PRJNA1070048 with BioProject ID PRJNA1070048.
References
Atteya AK, El-Serafy RS, El-Zabalawy KM, Elhakem A, Genaidy EA (2022) Brassinolide maximized the fruit and oil yield, induced the secondary metabolites, and stimulated linoleic acid synthesis of Opuntia ficus-indica oil. Horticulturae 8(5):452. https://doi.org/10.3390/plants12051133
Bajguz A, Tretyn A (2003) The chemical characteristic and distribution of brassinosteroids in plants. Phytochemistry 62:1027–1046. https://doi.org/10.1016/s0031-9422(02)00656-8
Bao AL, Burritt DJ, Chen HF, Zhou XA, Cao D, Tran LP (2019a) The CRISPR/Cas9 system and its applications in crop genome editing. Crit Rev Biotechnol 39:321–336. https://doi.org/10.1080/07388551.2018.1554621
Bao AL, Chen HF, Chen LM, Chen SL, Hao QN, Guo W, Qiu DZ, Shan ZH, Yang ZL, Yuan SL, Zhang CJ, Zhang XJ, Liu BH, Kong FJ, Li X, Zhou XA, Tran LSP, Cao D (2019b) CRISPR/Cas9-mediated targeted mutagenesis of GmSPL9 genes alters plant architecture in soybean. BMC Plant Biol 19:131. https://doi.org/10.1186/s12870-019-1746-6
Cao D, Li Y, Wang JL, Nan HY, Wang YN, Lu SJ, Jiang Q, Li XM, Shi DN, Fang C, Yuan XH, Zhao XH, Li X, Liu BH, Kong FJ (2015) GmmiR156b overexpression delays flowering time in soybean. Plant Mol Biol 89:353–363. https://doi.org/10.1007/s11103-015-0371-5
Chen LM, Yang HL, Fang YS, Guo W, Chen HF, Zhang XJ, Dai WJ, Chen SL, Hao QN, Yuan SL, Zhang CJ, Huang Y, Shan ZH, Yang ZL, Qiu DZ, Liu XR, Tran LSP, Zhou XA, Cao D (2020) Overexpression of GmMYB14 improves high-density yield and drought tolerance of soybean through regulating plant architecture mediated by the brassinosteroid pathway. Plant Biotechnol J 19:702–716. https://doi.org/10.1111/pbi.13496
Cheng L, Li M, Min W, Wang M, Chen R, Wang W (2021) Optimal brassinosteroid levels are required for soybean growth and mineral nutrient homeostasis. Int J Mol Sci 22:8400. https://doi.org/10.3390/ijms22168400
Deslauriers SD, Larsen PB (2010) FERONIA is a key modulator of brassinosteroid and ethylene responsiveness in Arabidopsis hypocotyls. Mol Plant 3:626–640. https://doi.org/10.1093/mp/ssq015
Dorion S, Ouellet JC, Rivoal J (2021) Glutathione metabolism in plants under stress: beyond reactive oxygen species detoxification. Metabolites 11:641. https://doi.org/10.3390/metabo11090641
Fang C, Du HP, Wang LS, Liu BH, Kong FJ (2023) Mechanisms underlying key agronomic traits and implications for molecular breeding in soybean. J Genet Genomics. https://doi.org/10.1016/j.jgg.2023.09.004
Gao JS, Yang SX, Cheng W, Fu YF, Leng JT, Yuan XH, Jiang N, Ma JX, Feng XZ (2017) GmILPA1, encoding an APC8-like protein, controls leaf petiole angle in soybean. Plant Physiol 174:1167–1176. https://doi.org/10.1104/pp.16.00074
Han C, Wang LY, Lyu JY, Shi W, Yao LM, Fan M, Bai M-Y (2023) Brassinosteroid signaling and molecular crosstalk with nutrients in plants. J Genet Genomics 50:541–553. https://doi.org/10.1016/j.jgg.2023.03.004
Hong Z, Ueguchi-Tanaka M, Fujioka S, Takatsuto S, Yoshida S, Hasegawa Y, Ashikari M, Kitano H, Matsuoka M (2005) The rice brassinosteroid-deficient dwarf2 mutant, defective in the rice homolog of Arabidopsis DIMINUTO/DWARF1, is rescued by the endogenously accumulated alternative bioactive brassinosteroid, dolichosterone. Plant Cell 17:2243–2254. https://doi.org/10.1105/tpc.105.030973
Huo WG, Li BD, Kuang JB, He PG, Xu ZH, Wang JX (2018) Functional characterization of the steroid reductase genes GmDET2a and GmDET2b from glycine max. Int J Mol Sci 19:726. https://doi.org/10.3390/ijms19030726
Klahre U, Noguchi T, Fujioka S, Takatsuto S, Yokota T, Nomura T, Yoshida S, Chua NH (1998) The Arabidopsis DIMINUTO/DWARF1 gene encodes a protein involved in steroid synthesis. Plant Cell 10:1677–1690. https://doi.org/10.1105/tpc.10.10.1677
Knoch E, Sugawara S, Mori T, Poulsen C, Fukushima A, Harholt J, Fujimoto Y, Umemoto N, Saito K (2018) Third DWF1 paralog in Solanaceae, sterol Δ 24 -isomerase, branches withanolide biosynthesis from the general phytosterol pathway. Proc Natl Acad Sci 115:34. https://doi.org/10.1073/pnas.1807482115
Liu YF, Zhang DJ, Ping JQ, Li S, Chen ZX, Ma JX (2016) Innovation of a regulatory mechanism modulating semi-determinate stem growth through artificial selection in soybean. PLoS Genet 12(1):e1005818. https://doi.org/10.1371/journal.pgen.1005818
Liu X, Yang H, Wang Y, Zhu Z, Zhang W, Li J (2020) Comparative transcriptomic analysis to identify brassinosteroid response genes. Plant Physiol 184(2):1072–1082. https://doi.org/10.1104/pp.20.00386
Ma XL, Zhang QY, Zhu QL, Liu W, Chen Y, Qiu R, Wang B, Yang ZF, Li HY, Lin YR, Xie YY, Shen RX, Chen SF, Wang Z, Chen YL, Guo JX, Chen LT, Zhao XC, Dong ZC, Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8:1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Miller CN, Harper AL, Trick M, Wellner N, Werner P, Waldron KW, Bancroft I (2018) Dissecting the complex regulation of lodging resistance in Brassica napus. Mol Breeding 38:30. https://doi.org/10.1007/s11032-018-0781-6
Nelson BK, Cai X, Nebenführ A (2007) A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J 51:1126–1136. https://doi.org/10.1111/j.1365-313X.2007.03212.x
Ohashi S, Kurita H, Takahashi Y, Nagasuga K, Nagaya Y, Umezaki T (2021) Varietal difference of soybean plant type focus on petiole. Plant Prod Sci 24(1):103–108. https://doi.org/10.1080/1343943X.2020.1801349
Peng SN, Tao P, Xu F, Wu AP, Huo WG, Wang JX (2016) Functional characterization of soybean Glyma04g39610 as a brassinosteroid receptor gene and evolutionary analysis of soybean brassinosteroid receptors. Int J Mol Sci 17:897. https://doi.org/10.3390/ijms17060897
Ping JQ, Liu YF, Sun LJ, Zhao MX, Li YH, She MY, Sui Y, Lin F, Liu XD, Tang ZX, Nguyen H, Tian ZX, Qiu LJ, Nelson RL, Clemente TE, Specht JE, Ma JX (2014) Dt2 is a gain-of-function MADS-domain factor gene that specifies semideterminacy in soybean. Plant Cell 26:2831–2842. https://doi.org/10.1105/tpc.114.126938
Rai GK, Kumar P, Choudhary SM, Singh H, Adab K, Kosser R et al (2023) Antioxidant potential of glutathione and crosstalk with phytohormones in enhancing abiotic stress tolerance in crop plants. Plants 12(5):1133. https://doi.org/10.3390/plants12051133
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Shimada Y, Goda H, Nakamura A, Takatsuto S, Fujioka S, Yoshida S (2003) Organ-specific expression of brassinosteroid-biosynthetic genes and distribution of endogenous brassinosteroids in Arabidopsis. Plant Physiol 131:287–297. https://doi.org/10.1104/pp.013029
Song L, Chen W, Wang B, Yao Q-M, Valliyodan B, Bai M-Y, Zhao M-Z, Ye H, Wang Z-Y, Nguyen HT (2019a) GmBZL3 acts as a major BR signaling regulator through crosstalk with multiple pathways in Glycine max. BMC Plant Biol 19:86. https://doi.org/10.1186/s12870-019-1677-2
Song L, Chen W, Yao Q, Guo B, Valliyodan B, Wang Z, Nguyen HT (2019b) Genome-wide transcriptional profiling for elucidating the effects of brassinosteroids on Glycine max during early vegetative development. Sci Rep 9(1):16085. https://doi.org/10.1038/s41598-019-52599-3
Song L, Liu J, Cao B, Liu B, Zhang X, Chen Z, Dong C, Liu X, Zhang Z, Wang W, Chai L, Liu J, Zhu J, Cui S, He F, Peng H, Hu Z, Su Z, Guo W, Xin M, Yao Y, Yan Y, Song Y, Bai G, Sun Q, Ni Z (2023) Reducing brassinosteroid signalling enhances grain yield in semi-dwarf wheat. Nature 617:118–124. https://doi.org/10.1038/s41586-023-06023-6
Sparkes IA, Runions J, Kearns A, Hawes C (2006) Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat Protoc 1:2019–2025. https://doi.org/10.1038/nprot.2006.286
Sun Z, Su C, Yun J, Jiang Q, Wang L, Wang Y, Cao D, Zhao F, Zhao Q, Zhang M, Zhou B, Zhang L, Kong F, Liu B, Tong Y, Li X (2019) Genetic improvement of the shoot architecture and yield in soya bean plants via the manipulation of GmmiR156b. Plant Biotechnol J 17:50–62. https://doi.org/10.1111/pbi.12946
Tanaka K, Asami T, Yoshida S, Nakamura Y, Matsuo T, Okamoto S (2005) Brassinosteroid homeostasis in Arabidopsis is ensured by feedback expressions of multiple genes involved in its metabolism. Plant Physiol 138:1117–1125. https://doi.org/10.1104/pp.104.058040
Tao Y, Zheng J, Xu Z, Zhang X, Zhang K, Wang G (2004) Functional analysis of ZmDWF1, a maize homolog of the Arabidopsis brassinosteroids biosynthetic DWF1/DIM gene. Plant Sci 167:743–751. https://doi.org/10.1016/j.plantsci.2004.05.012
Tian ZX, Wang XB, Lee R, Li YH, Specht JE, Nelson RL, McClean PE, Qiu LJ, Ma JX (2010) Artificial selection for determinate growth habit in soybean. Proc Natl Acad Sci USA 107:8563–8568. https://doi.org/10.1073/pnas.1000088107
Tong H, Chu C (2018) Functional specificities of brassinosteroid and potential utilization for crop improvement. Trends Plant Sci 23:1016–1028. https://doi.org/10.1016/j.tplants.2018.08.007
Vriet C, Russinova E, Reuzeau C (2012) Boosting crop yields with plant steroids. Plant Cell 24:842–857. https://doi.org/10.1105/tpc.111.094912
Yang Q, Lin G, Lv H, Wang C, Yang Y, Liao H (2021) Environmental and genetic regulation of plant height in soybean. BMC Plant Biol 21:1–15. https://doi.org/10.1186/s12870-021-02836-7
Yin W, Dong N, Niu M, Zhang X, Li L, Liu J, Liu B, Tong H (2019) Brassinosteroid-regulated plant growth and development and gene expression in soybean. Crop J 7:411–418. https://doi.org/10.1016/j.cj.2018.10.003
Youn JH, Kim TW, Joo SH, Son SH, Roh J, Kim S, Kim TW, Kim SK (2018) Function and molecular regulation of DWARF1 as a C-24 reductase in brassinosteroid biosynthesis in Arabidopsis. J Exp Bot 69:1873–1886. https://doi.org/10.1093/jxb/ery038
Zhang Z, Finer JJ (2015) Soybean actin, heat shock protein, and ribosomal protein promoters direct tissue-specific transgene expression in transgenic soybean. In Vitro Cell Dev Biol—Plant 51:9–18. https://doi.org/10.1007/s11627-014-9656-7
Zhang Y, Zhang Y-J, Yang B-J, Yu X-X, Wang D, Zu S-H, Xue H-W, Lin W-H (2016) Functional characterization of GmBZL2 (AtBZR1 like gene) reveals the conserved BR signaling regulation in Glycine max. Sci Rep 6:31134. https://doi.org/10.1038/srep31134
Zheng X, Xiao Y, Tian Y, Yang S, Wang C (2020) PcDWF1, a pear brassinosteroid biosynthetic gene homologous to AtDWARF1, affected the vegetative and reproductive growth of plants. BMC Plant Biol 20:109. https://doi.org/10.1186/s12870-020-2323-8
Funding
The research was supported by The National Natural Science Foundation of China (32071926 and 32072087); the Agricultural Science and Technology Innovation Program of Chinese Academy of Agricultural Sciences (CAAS-OCRI-XKPY-202103).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: LL, WG, HC, Performed the experiments: HY, XX, XY, CD, SM, QZ. Analyzed the data: HY, XX, XY, LC. Contributed reagents/materials/analysis tools: HY, XX. Wrote the paper: XX, HY, WG. Revised the paper: LL, WG, DC. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no competing interests.
Additional information
Communicated by Haitao Shi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiang, X., Yang, H., Yuan, X. et al. CRISPR/Cas9-mediated editing of GmDWF1 brassinosteroid biosynthetic gene induces dwarfism in soybean. Plant Cell Rep 43, 116 (2024). https://doi.org/10.1007/s00299-024-03204-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00299-024-03204-z