Abstract
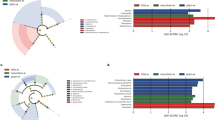
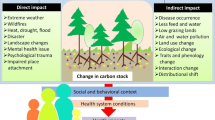
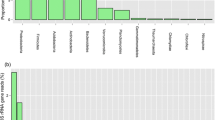
The presence of microbial communities in the rhizosphere of plants is an important determinant of plant health and soil organic matter composition. Plant species play significant roles in selecting the specific microbial communities that inhabit the root zone. However, till now, there is no solid information regarding the presence of specific plant-microbiome in the rhizosphere of many plants, especially under-exploited and under-researched species such as Kersting’s groundnut. This study assessed the effect of five Kersting’s groundnut landraces on the structure of microbial communities in rhizosphere of field-grown plants. The five tested Kersting’s groundnut landraces (Belane Mottled, Boli, Funsi, Puffeun and Heng Red Mottled) were found to exert a marked selective influence on bacteria associated with their rhizospheres, measured using 16S rDNA MiSeq illumina sequencing. Community differences in microbial composition and relative abundance were both significant. Numerous phyla in the rhizosphere were affected by the test landraces. Except for Belane mottled whose rhizospheres were dominated by Proteobacteria, the rhizosphere soils of the other landraces were dominated by Bacteroidetes. With the exception of landrace Puffeun which showed only Mesorhizobium in its rhizosphere, all the other test landraces revealed the presence of Bradyrhizobium and Rhizobium species of alpha Proteobacteria. Furthermore, the rhizosphere of all landraces were abundant in species of the indole-3-acetic–acid producing Sphingomonas and cellulose-degrading Fibrobacteres. The results of this study suggest that Kersting’s groundnut landraces can shape bacterial community composition in the rhizosphere via plant-related changes in the rhizosphere soil.







Similar content being viewed by others
References
Dzoyem JP, Kuete V, Eloff JN (2014) Biochemical parameters in toxicological studies in Africa: significance, principle of methods, data interpretation, and use in plant screenings. Toxicological Survey of African Medicinal Plants. Elsevier, Amsterdam, pp 659–715
Group AP (2009) An update of the angiosperm phylogeny group classification for the orders and families of flowering plants: APG III. Bot J Linn Soc 161:105–121
Magbagbeola JAO, Adetoso JA, Owolabi OA (2010) Neglected and underutilized species (NUS): a panacea for community focused development to poverty alleviation/poverty reduction in Nigeria. J Econ Int Financ 2:208
Pasquet RS, Mergeai G, Baudoin J-P (2002) Genetic diversity of the African geocarpic legume Kersting’s groundnut, Macrotyloma geocarpum (Tribe Phaseoleae: Fabaceae). Biochem Syst Ecol 30:943–952. https://doi.org/10.1016/S0305-1978(02)00040-6
Bampuori AH (2007) Effect of traditional farming practices on the yield of indigenous Kersting’s groundnut (Macrotyloma geocarpum Harms) crop in the upper west region of Ghana. J Dev Sustain Agric 2:128–144
Assogba P, Ewedje EEBK, Dansi A et al (2016) Indigenous knowledge and agro-morphological evaluation of the minor crop Kersting’s groundnut [Macrotyloma geocarpum (Harms) Marechal et Baudet] cultivars of Benin. Genet Resour Crop Evolut 63:513–529
Ajayi OB, Oyetayo FL (2009) Potentials of Kerstingiella geocarpa as a health food. J Med Food 12:184–187
Ayenan MAT, Ezin VA (2016) Potential of Kersting’s groundnut [Macrotyloma geocarpum (Harms) Marechal & Baudet] and prospects for its promotion. Agric Food Secur 5:10
Adu-Gyamfi R, Dzomeku IK, Lardi J et al (2012) Evaluation of growth and yield potential of genotypes of Kersting’s groundnut (Macrotyloma geocarpum harms) in Northern Ghana. Int Res J Agric Sci Soil Sci 2:509–515
Dansi A, Vodouhè R, Azokpota P et al (2012) Diversity of the neglected and underutilized crop species of importance in Benin. Sci World J. https://doi.org/10.1100/2012/932947
Mohammed M, Jaiswal SK, Sowley ENK, Ahiabor BDK (2018) Symbiotic N2 fixation and grain yield of endangered Kersting’s groundnut landraces in response to soil and plant associated Bradyrhizobium inoculation to promote ecological resource-use efficiency. Front Microbiol 9:2105
Amuti K (1980) Geocarpa groundnut (Kerstingiella geocarpa) in Ghana. Econ Bot 34:358–361
Dakora FD (1998) Nodule function in symbiotic Bambara groundnut (Vigna subterranea L.) and Kersting’s bean (Macrotyloma geocarpum L.) is tolerant of nitrate in the root medium. Ann Bot 82:687–690
Mohammed M, Jaiswal SK, Dakora FD (2019) Insights into the phylogeny, nodule functioning and biogeographic distribution of microsymbionts nodulating the orphan Kersting’s groundnut [Macrotyloma geocarpum (Harms) Marechal & Baudet] in African soils. Appl Environ Microbiol 85(11):e00342
Makoi J, Ndakidemi P (2007) Biological, ecological and agronomic significance of plant phenolic compounds in rhizosphere of the symbiotic legumes. African J Biotechnol 6(12):1358
Hassan S, Mathesius U (2012) The role of flavonoids in root-rhizosphere signalling: opportunities and challenges for improving plant-microbe interactions. J Exp Bot 63:3429–3444
Dakora FD, Phillips DA (2002) Root exudates as mediators of mineral acquisition in low-nutrient environments. Plant Soil 245:35–47
Sugiyama A, Ueda Y, Zushi T et al (2014) Changes in the bacterial community of soybean rhizospheres during growth in the field. PLoS ONE 9:1–9
Carvalho FM, Souza RC, Barcellos FG et al (2010) Genomic and evolutionary comparisons of diazotrophic and pathogenic bacteria of the order Rhizobiales. BMC Microbiol 10:37
Trabelsi D, Mhamdi R (2013) Microbial inoculants and their impact on soil microbial communities: a review. Biomed Res Int. https://doi.org/10.1155/2013/863240
Kawasaki A, Watson ER, Kertesz MA (2012) Indirect effects of polycyclic aromatic hydrocarbon contamination on microbial communities in legume and grass rhizospheres. Plant Soil 358:169–182
Bloemberg GV, Lugtenberg BJJ (2001) Molecular basis of plant growth promotion and biocontrol by rhizobacteria. Curr Opin Plant Biol 4:343–350
Philippot L, Raaijmakers JM, Lemanceau P, van der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789
Kuzyakov Y, Domanski G et al (2000) Carbon input by plants into the soil. Review. J Plant Nutr Soil Sci 163:421–431
Mendes R, Kruijt M, De Bruijn I et al (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100
Kuzyakov Y (2002) Factors affecting rhizosphere priming effects. J Plant Nutr Soil Sci 165:382–396
Bhattacharjee RB, Singh A, Mukhopadhyay SN (2008) Use of nitrogen-fixing bacteria as biofertiliser for non-legumes: prospects and challenges. Appl Microbiol Biotechnol 80:199–209
Micallef SA, Shiaris MP, Colón-Carmona A (2009) Influence of Arabidopsis thaliana accessions on rhizobacterial communities and natural variation in root exudates. J Exp Bot 60:1729–1742
Reynolds HL, Packer A, Bever JD, Clay K (2003) Grassroots ecology: plant–microbe–soil interactions as drivers of plant community structure and dynamics. Ecology 84:2281–2291
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev 68:669–685
Roesch LFW, Fulthorpe RR, Riva A et al (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1:283
Badii BK, Adarkwah C, Obeng-Ofori D, Ulrichs C (2014) Efficacy of diatomaceous earth formulations against Callosobruchus maculatus (F.) (Coleoptera: Bruchidae) in Kersting’s groundnut (Macrotyloma geocarpum Harms): influence of dosage rate and relative humidity. J Pest Sci 87:285–294
Badii KB, Asante SK, Bayorbor TB (2011) Susceptibility of some kersting’s groundnut landrace cultivars to infestation by Callosobruchus maculatus (fab.)[Coleoptera: Bruchidae]. J Sci Technol 31:11–20
Margulies M, Egholm M, Altman WE et al (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376
Schöler A, Jacquiod S, Vestergaard G et al (2017) Analysis of soil microbial communities based on amplicon sequencing of marker genes. Biol Fertil Soils 53:485–489
Tsamo AT, Mohammed M, Ndibewu PP, Dakora FD (2019) Identification and quantification of anthocyanins in seeds of Kersting’s groundnut [Macrotyloma geocarpum (Harms) Marechal & Baudet] landraces of varying seed coat pigmentation. J Food Meas Charact. https://doi.org/10.1007/s11694-019-00150-3
Vestergaard G, Schulz S, Schöler A, Schloter M (2017) Making big data smart—how to use metagenomics to understand soil quality. Biol Fertil Soils 53:479–484
Klindworth A, Pruesse E, Schweer T et al (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1–e1
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Method 7:335
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Aslam Z, Yasir M, Khaliq A et al (2010) Mini review: too much bacteria still unculturable. Bioscience 45:600–609
Turner TR, Ramakrishnan K, Walshaw J et al (2013) Comparative metatranscriptomics reveals kingdom level changes in the rhizosphere microbiome of plants. ISME J 7:2248–2258. https://doi.org/10.1038/ismej.2013.119
İnceoğlu Ö, Al-Soud WA, Salles JF et al (2011) Comparative analysis of bacterial communities in a potato field as determined by pyrosequencing. PLoS ONE 6:e23321
Lundberg DS, Lebeis SL, Paredes SH et al (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90. https://doi.org/10.1038/nature11237
Urich T, Lanzén A, Qi J et al (2008) Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome. PLoS ONE 3:e2527
Lu Y, Rosencrantz D, Liesack W, Conrad R (2006) Structure and activity of bacterial community inhabiting rice roots and the rhizosphere. Environ Microbiol 8:1351–1360
Hunt S, Layzell DB (1993) Gas exchange of legume nodules and the regulation of nitrogenase activity. Annu Rev Plant Biol 44:483–511
Muleta D, Assefa F, Börjesson E, Granhall U (2013) Phosphate-solubilising rhizobacteria associated with Coffea arabica L. in natural coffee forests of southwestern Ethiopia. J Saud Soc Agric Sci 12:73–84
Dardanelli MS, de Córdoba FJF, Estévez J et al (2012) Changes in flavonoids secreted by Phaseolus vulgaris roots in the presence of salt and the plant growth-promoting rhizobacterium Chryseobacterium balustinum. Appl Soil Ecol 57:31–38
Miethling R, Ahrends K, Tebbe CC (2003) Structural differences in the rhizosphere communities of legumes are not equally reflected in community-level physiological profiles. Soil Biol Biochem 35:1405–1410
Singh AV, Chandra R, Goel R (2013) Phosphate solubilization by Chryseobacterium sp. and their combined effect with N and P fertilizers on plant growth promotion. Arch Agron Soil Sci 59:641–651
Ransom-Jones E, Jones DL, McCarthy AJ, McDonald JE (2012) The fibrobacteres: an important phylum of cellulose-degrading bacteria. Microb Ecol 63:267–281
Dennis PG, Miller AJ, Hirsch PR (2010) Are root exudates more important than other sources of rhizodeposits in structuring rhizosphere bacterial communities? FEMS Microbiol Ecol 72:313–327
Galbally IE, Kirstine W (2002) The production of methanol by flowering plants and the global cycle of methanol. J Atmos Chem 43:195–229
Mazzola M, Funnell DL, Raaijmakers JM (2004) Wheat cultivar-specific selection of 2,4-diacetylphloroglucinol-producing fluorescent Pseudomonas species from resident soil populations. Microb Ecol 48:338–348
Dalmastri C, Chiarini L, Cantale C et al (1999) Soil type and maize cultivar affect the genetic diversity of maize root-associated Burkholderia cepacia populations. Microb Ecol 38:273–284
Badri DV, Chaparro JM, Zhang R et al (2013) Application of natural blends of phytochemicals derived from the root exudates of Arabidopsis to the soil reveal that phenolic-related compounds predominantly modulate the soil microbiome. J Biol Chem 288:4502–4512
Bais HP, Weir TL, Perry LG et al (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol 57:233–266
Bertin C, Yang X, Weston LA (2003) The role of root exudates and allelochemicals in the rhizosphere. Plant Soil 256:67–83
Broeckling CD, Broz AK, Bergelson J et al (2008) Root exudates regulate soil fungal community composition and diversity. Appl Environ Microbiol 74:738–744
Shi S, Richardson AE, O’Callaghan M et al (2011) Effects of selected root exudate components on soil bacterial communities. FEMS Microbiol Ecol 77:600–610
Gu Y, Wei Z, Wang X et al (2016) Pathogen invasion indirectly changes the composition of soil microbiome via shifts in root exudation profile. Biol Fertil Soils 52:997–1005
Czarnota MA, Rimando AM, Weston LA (2003) Evaluation of root exudates of seven sorghum accessions. J Chem Ecol 29:2073–2083. https://doi.org/10.1023/A:1025634402071
Warembourg FR, Roumet C, Lafont F (2003) Differences in rhizosphere carbon-partitioning among plant species of different families. Plant Soil 256:347–357. https://doi.org/10.1023/A:1026147622800
Acknowledgements
This work was supported with grants from the South African Department of Science and Technology, the Tshwane University of Technology, the National Research Foundation in Pretoria, and the South African Research Chair in Agrochemurgy and Plant Symbioses.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jaiswal, S.K., Mohammed, M. & Dakora, F.D. Microbial community structure in the rhizosphere of the orphan legume Kersting’s groundnut [Macrotyloma geocarpum (Harms) Marechal & Baudet]. Mol Biol Rep 46, 4471–4481 (2019). https://doi.org/10.1007/s11033-019-04902-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04902-8